Here
is a very simple paper from Nature. Sometimes
simple equals great. Not this time, though. No wonder it
took 1.5 year to have it published.
This
is a study about Aryl Hydrocarbon receptor (AhR). As I have discussed
in earlier posts, AhR senses environmental pollutants and toxins,
like p-dioxin. More
importantly, however, AhR senses natural, endogenously-generated
molecules like amino acid tryptophan degradation products. Upon
ligand engagement, AhR activates detoxifying enzymes.
Here
(1),
the authors tested the hypothesis the AhR can detect
pathogen-associated molecules. Using in
silico modeling
they identify several pigmented
virulence factors derived
from P. aeruginosa (phenazine)
and M. tuberculosis (phthiocol)
that could bind AhR.
To
verify this finding in vitro the authors used
luciferase reporter assay with human macrophage cell line (THP-1)
transfected with AhR responsive elements. Indeed, physiological
concentration of phenazine or phthiocol could activate AhR.
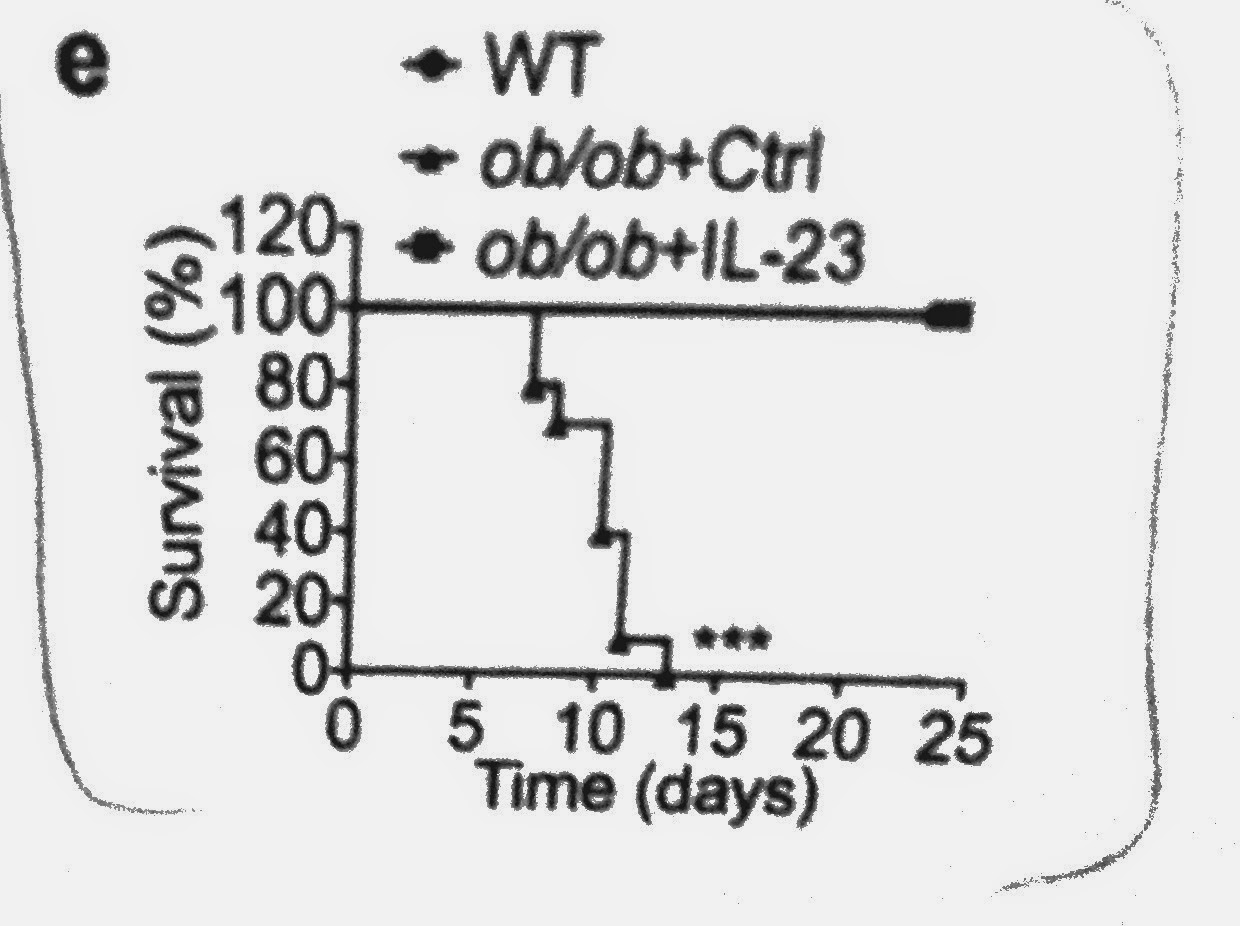
To
test this observation in vivo, the authors
infected AhR-KO mice with P. aeruginosa. Compared to
wild-type control, AhR-KO mice succumbed to infection more rapidly.
This was associated with more tissue damage and less neutrophil
infiltration.
Bone
marrow chimera experiment indicated that both
haematopoietic and non-haematopoietic cell expression of AhR were
necessary for resistance to P. aeruginosa infection.
In
addition, the authors showed that resistance to M. tuberculosis (both
at low or high dose) was also influenced by absence of AhR signaling.
In summary, the authors
postulate that AhR can sense infection-derived molecules and
contributes to host defense.
While
this model is very simple, there are few weaknesses in this paper.
Since AhR function is quite broad, to show its relevance in
infection-derived molecule detection, the authors should have used
AhR-ligand deficient P. aeruginosa or
M. tuberculosis, for example, mutant strain PA14 phz ½.
However, the authors did not used it. In
absence of such experiment, however, the authors conclusion regarding
pattern-recognition receptor function of AhR is questionable.